Part 3: Mutating Proteins and MD Setup
Getting Started
To start, we must first download the files that you will need for this section of the workshop. You can download the files here (mutation.tgz).
Change into the directory in which you have downloaded mutation.tgz and unpack it using the command
tar -zxvf mutation.tgzThis will create a directory called ‘mutation’. Change into this directory by typing;
cd mutationIf you look in this directory you will see the following files;
:-> ls
1NNC.pdb ZAN.frcmod ZAN.prepin add_atoms.in add_water.in ca.frcmod ca.prep equilconfig heatconfig mdconfig minconfigThere are a lot of files. The first one we will look at is ‘1NNC.pdb’. This is a PDB file of zanamivir bound to the wild type form of H7N9 neuraminidase. You can look at this file by typing;
vmd 1NNC.pdb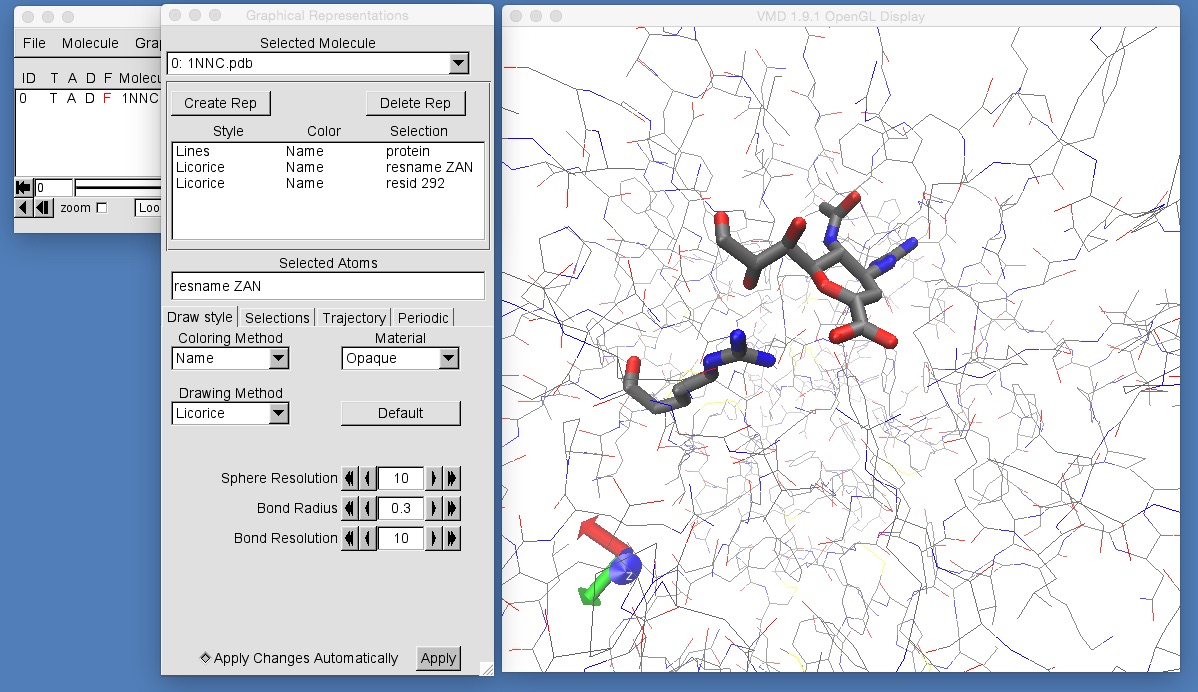
You will see that the protein and zanamivir are missing hydrogen atoms, and that this is not in a box of water. You can view zanamivir by creating a graphical representation using selection string “resname ZAN”. You can view arginine 292 by creating a graphical representation using selection string “resid 292”.