Part 1: Molecular Visualisation
Selected Atoms
The “Selected Atoms” box provides a way to select only a subset of atoms to draw. The ‘h7n9.pdb’ file that you have loaded actually contains more than just the H7N9 neuraminidase protein. It also contains a single oseltamivir drug molecule that is bound in the active site.
- To view this molecule, click “Create Rep” to create a new representation.
- Then, in the “Selected Atoms” box, change “All” to read “resname OSE”.
- Then change the coloring method to “Name” and the drawing method to “Licorice”.
If you do this, you should see the following;
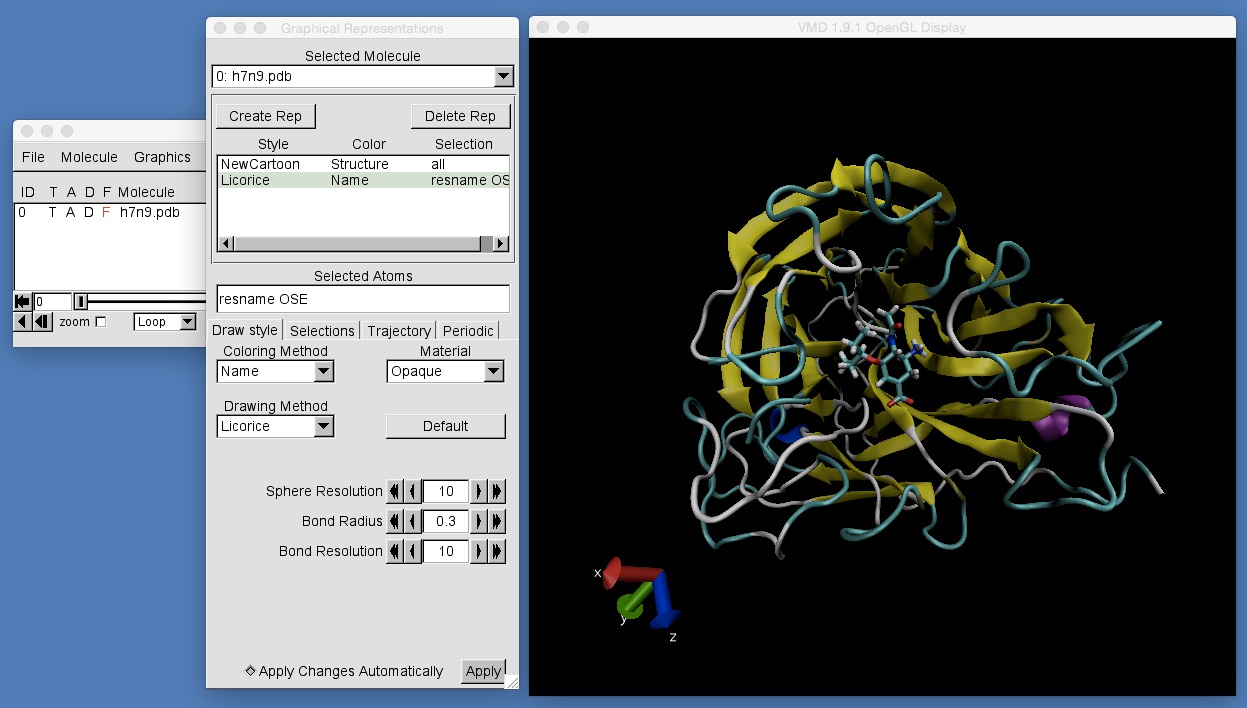
To help with categorising, we arrange atoms in molecules into residues. The “resname OSE” selection string tells VMD to select only the atoms that are part of the residue called “OSE”.
How do we know that the atoms in oseltamivir are in the residue called “OSE”? There are two ways to know. We can either take a look in the original PDB file and look for the lines that describe these atoms, e.g. on line 5940 of h7n9.pdb we find;
ATOM 5935 HD23 LEU 388 20.452 36.384 64.467 0.00 0.00
ATOM 5936 C LEU 388 18.189 32.762 65.557 0.00 0.00
ATOM 5937 O LEU 388 17.033 32.579 65.892 0.00 0.00
ATOM 5938 OXT LEU 388 19.161 32.097 65.932 0.00 0.00
TER 5939 LEU 388
ATOM 5940 O3 OSE 389 39.883 39.200 43.708 0.00 0.00
ATOM 5941 C6 OSE 389 39.404 39.770 42.740 0.00 0.00
ATOM 5942 C7 OSE 389 39.561 39.252 41.385 0.00 0.00
ATOM 5943 H5 OSE 389 39.590 38.162 41.442 0.00 0.00
ATOM 5944 H6 OSE 389 40.543 39.507 40.982 0.00 0.00
ATOM 5945 H7 OSE 389 38.744 39.520 40.712 0.00 0.00
ATOM 5946 N1 OSE 389 38.466 40.721 42.797 0.00 0.00
ATOM 5947 H23 OSE 389 38.012 40.929 41.920 0.00 0.00The “TER” is used in a PDB file to divide a file into multiple molecules. This “TER” is dividing the protein from the drug molecule, and so we can see that the atoms of the drug are placed into the residue called “OSE” (the fourth column in each line).
An easier way to find that the drug is called “OSE” is to use the “Selections” tab in VMD’s “Graphical Representations” window. In the “Graphical Representations” window click on “Selections” (one of four tabs, “Draw Style”, “Selections”, “Trajectory” and “Periodic”), and you will see the following.
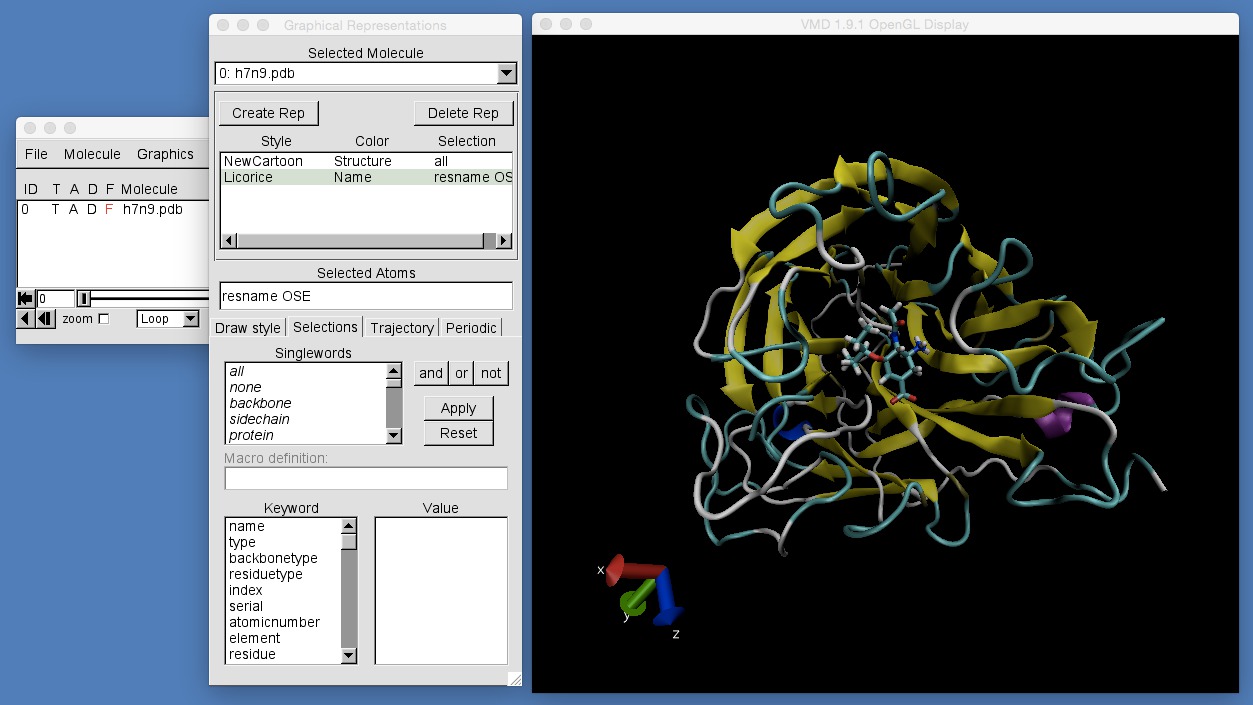
This tab shows you all of the selection strings that are possible. The first box contains the single-word selection strings, e.g.
- “all” - selects all of the atoms
- “none” - selects no atoms
- “backbone” - selects only protein backbone atoms
- “sidechain” - selects only protein sidechain atoms
- “protein” - selects only protein atoms
- “water” - selects only water atoms
- “alpha_helix” - selects only atoms part of an alpha helix
The second box contains the “keyword” type selections. One of these keywords is “resname” which means to select atoms by the name of their residue. Click on “resname”. This will show, in the right-hand box, all of the residue names of the loaded molecule. Scroll down and you will see that “OSE” is one of these names, e.g.
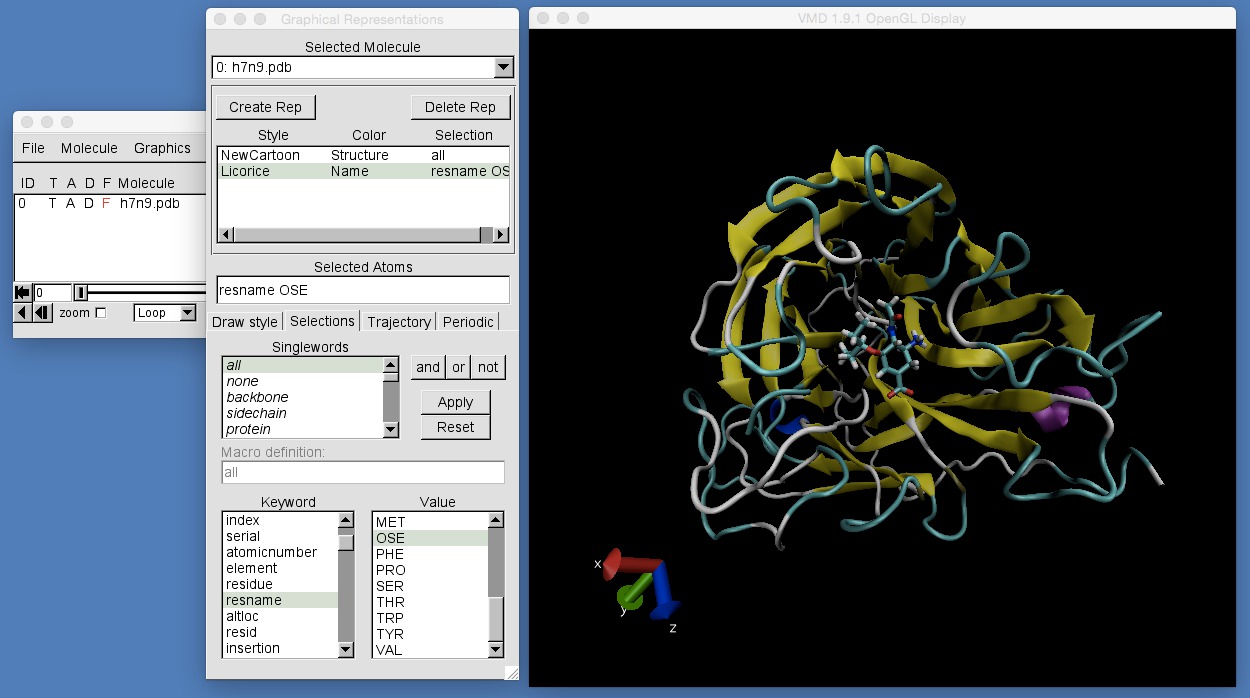
In addition to selecting atoms by the name of their residue, you can also use these “keyword” selections to select atoms by name (using the “name” keyword), by residue number (using the “resid” keyword) and by residue indices (using the “residue” keyword).
To choose a selection, edit the string in the “Selected Atoms” box so that it contains the text that you want, e.g. to select all atoms in residue 350, change the “Selected Atoms” string to read;
residue 350You should find that this selects one of the residues in the protein.
Play around with different selections and representations. Try creating different representations for the alpha helices, protein sidechains and oseltamivir.